Results from the extensive computation analysis of the 1029 sequenced genomes from India carried out by CSIR constituent labs, CSIR-Institute of Genomics and Integrative Biology (IGIB), Delhi and CSIR-Centre for Cellular and Molecular Biology (CCMB), Hyderabad were published in the scientific journal, Nucleic Acid Research, earlier this week. The allele frequencies of the genetic variants generated in the study are available on the IndiGenomesdatabase accessible at http://clingen.igib.res.in/indigen.
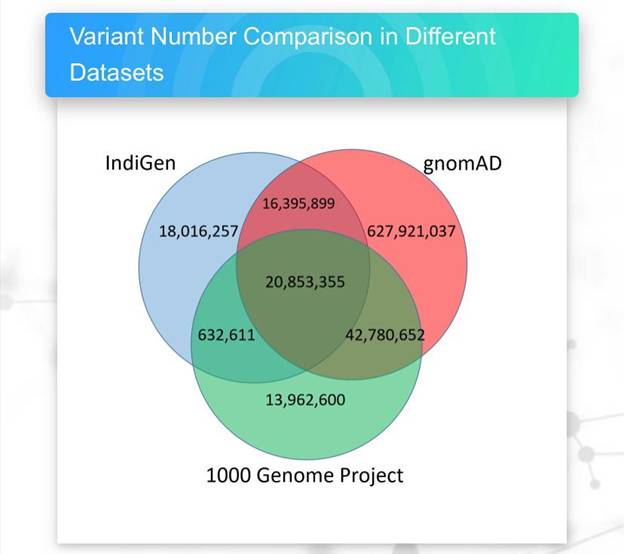
The analysis led to the identification of 55,898,122 single nucleotide variants in the India genome dataset. Comparisons with the global genome datasets revealed that 18,016,257 (32.23%) variants were unique and found only in the samples sequenced from India. This emphasizes the need for an India centric population genomic initiative.

India is the second largest country in terms of population density with more than 1.3 billion individuals encompassing 17% of the world population. Despite having this rich genetic diversity, India has been under-represented in global genome studies. Further, the population architecture of India has resulted in high prevalence of recessive alleles. In the absence of large-scale whole genome studies from India, these population-specific genetic variants are not adequately captured and catalogued in global medical literature.
In order to fill the gap of whole genome sequences from different populations in India, CSIR initiated the IndiGen Program in April 2019. Under this program, the whole genome sequencing of 1029 self-declared healthy Indians drawn from across the country has been completed. This has enabled benchmarking the scalability of genome sequencing at population scale in a defined timeline. The Minister of S&T, ES and MoHFW, Dr Harsh Vardhan announced the completion of the IndiGen sequence generation efforts on 25th October 2019.
The current IndiGenomes data resource provides a compendium of genetic variants representing the contemporary Indian population with an objective to classify variants involved in mendelian disorders and improve precision medicine outcomes.
The resource can also enable the identification of markers for carrier screening, variations causing genetic diseases, prevention of adverse events and provide better diagnosis and optimal therapy through mining data of clinically actionable pharmacogenetic variants. The phased data will allow researchers to build Indian-specific reference genome dataset and efficiently impute haplotype information. This resource can provide useful insights for clinicians and researchers in comprehending genetics not only at the population level but at the individual level.
The resource is widely accessible to the researchers and clinicians in India and abroad. There have been over ∼200,000 pageviews on the IndiGenomes web page from users spanning 27 countries demonstrating the uniqueness of the resource.
DG CSIR, Dr Shekhar C Mande, complimented the teams from CSIR-IGIB and CSIR-CCMB who undertook the IndiGen program not only leading to the development of this valuable resource, but agumenting capability in genomics and understanding of India’s genomic diversity strengthening the country’s response to the current and future pandemics.
[More information about the CSIR IndiGen Program may be obtained from:
Director, CSIR- Institute of Genomics and Integrative Biology, Delhi, India. director@igib.res.in
Director, CSIR- Centre of Cellular and Molecular Biology, Hyderabad, India. director@ccmb.res.in]
*****
NB/KGS/(CSIR Release)

